Aaron Puri
Assistant Professor of Chemistry
Chemical Ecology, Drug Discovery, Microbiome, Natural Products, Microbiology, Biosynthesis, Secondary Metabolites

Biological Chemistry Program
Education
B.S. University of Chicago
Ph.D. Stanford University School of Medicine
Research
Chemical Ecology
Groups of bacteria are responsible for many important processes on earth, from human health to carbon cycling. While a great deal of progress has been made identifying the bacterial species that are involved in these activities, in most cases how these bacteria actually interact remains a mystery. We are interested in determining the molecular details of how these bacteria interact with each other and their environment.
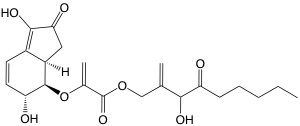
Natural Product Discovery and Biosynthesis
Natural products form the basis of many compounds essential to medicine and agriculture. Therefore, there is a constant demand for new sources of these molecules. The number of sequenced bacterial genomes has exploded in recent years, revealing a large untapped source of novel biosynthetic potential in species not traditionally relied upon for natural product discovery. We are interested in discovering new natural products with therapeutic potential from underexplored bacterial species. We are also interested in determing how these compounds are biosynthesized, in order to discover new chemistry that can be used to make novel compounds from renewable feedstocks in the future.

Regulation of Biosynthetic Gene Clusters
Bacteria often regulate the production of the natural products they make because they are energetically expensive. We are interested in determining how theses clusters are regulated, including how to turn on ones that are off in the laboratory (cryptic clusters) in order to increase the chance of discovering new compounds.
References
- Puri AW, Liu D, Schaefer AL, Yu Z, Pesesky MW, Greenberg EP, & Lidstrom ME. Interspecies chemical signaling in a methane-oxidizing bacterial community. Appl Environ Microbiol. 2019. Epub ahead of print.
- Puri AW*, Mevers E*, Ramadhar TR*, Petras D, Liu D, Piel J, Dorrestein PC, Greenberg EP, Lidstrom
ME, & Clardy J. Tundrenone: An Atypical Secondary Metabolite from Bacteria with Highly
Restricted Primary Metabolism. J Am Chem Soc. 2018. 140(6): 2002-2006. https://doi.org/10.1021/jacs.7b12240
*Equal contribution. - Puri AW, Schaefer AL, Fu Y, Beck DA, Greenberg EP, & Lidstrom ME. Quorum Sensing in a Methane-Oxidizing Bacterium. J Bacteriol. 2017. 199(5) pii: e00773-16. https://doi.org/10.1128/JB.00773-16
- Bender KO, Garland M, Ferreyra JA, Hryckowian AJ, Child MA, Puri AW, Solow-Cordero DE, Higginbottom SK, Segal E, Banaei N, Shen A, Sonnenburg JL, & Bogyo M. A small-molecule antivirulence agent for treating Clostridium difficile infection. Sci Transl Med. 2015. 7(306): 306ra148. https://doi.org/10.1126/scitranslmed.aac9103
- Puri AW, Owen S, Chu F, Chavkin T, Beck DA, Kalyuzhnaya MG, & Lidstrom ME. Genetic tools for the industrially promising methanotroph Methylomicrobium buryatense. Appl Environ Microbiol. 2015. 81(5): 1775-81. https://doi.org/10.1128/AEM.03795-14
- Xiao J, Broz P, Puri AW, Deu E, Morell M, Monack DM, & Bogyo M. A coupled protein and probe engineering approach for selective inhibition and activity-based probe labeling of the caspases. J Am Chem Soc. 2013. 135(24): 9130-8. https://doi.org/10.1021/ja403521u
- Puri AW, Broz P, Shen A, Monack DM, & Bogyo M. Caspase-1 activity is required to bypass macrophage apoptosis upon Salmonella infection. Nat Chem Biol. 2012. 8(9): 745-7. https://doi.org/10.1038/nchembio.1023
- Puri AW, Lupardus PJ, Deu E, Albrow VE, Garcia KC, Bogyo M, & Shen A. Rational design of inhibitors and activity-based probes targeting Clostridium difficile virulence factor TcdB. Chem Biol. 2010. 17(11): 1201-11. https://doi.org/10.1016/j.chembiol.2010.09.011
